Finalising a Structure
Before Writing a Final CIF
The bonds to hydrogen atoms are not automatically included in the CIF. Type >>htab in the command-line – this will print details of any hydrogen bonds that are found on the screen and include appropriate instructions relating to them into the refinement mode. You then need to refine the structure again to get these values in the CIF. Note that htab takes two default parameters, type >>help htab to find out more about this.
If a full analysis of torsion angles is required, type >>editins or click ![]() to open the .ins file header. Type
to open the .ins file header. Type >>CONF on a separate line somewhere below the UNIT line.
There is also a tick-box in the refinement settings (Work|Refine|CONF,MORE -1,Bond$H,ACTA) that will add a collection of commands to ensure that the final CIF will contain all of the information required by IUCr journals.
Editing Report / CIF parameters
Under Work|Report you can type a name for the the crystal structure report (‘data’ block name in CIF terminology). What follows will allow you to enter and/or modify details relating to the crystal and data collection. If you have solved and refined your structure in the “standard location” i.e. a folder that contains lots of other information about your experiment then most (if not all) of these fields should already be filled.
- Collection: information on the persons involved in the structure the collection date etc.
- Crystal: name, colour, size, shape preparation details etc.
- Diffraction: details on the diffractometer, diffraction temperature, special details. Clicking on Definition File brings up a number of files containing standard information on specific diffractometers. These files are located in the Olex2 directory/util/SiteSpecific and can be adjusted or new ones created (use a copy of template .cif) to fit users' own diffractometers.
- Absorption Correction: details of the absorption correction type, details, Tmax and Tmin.
- Publication: details of CCDC numbers, authors and journals.
- Reference: details on the published structure
- Source Files: Olex2 will extract as much information from the files present in your structure folder as possible. To do this, it tries to locate various files that might contain useful information and will add this information to what it already knows about your structure. This tool allows you to see what these files are and if we have used a “wrong” file, you can point Olex2 in the right direction.
Generating a Report
Under Work|Report are a number of boxes giving options for formatting reports. Image allows the inclusion of an image in the report. Choose a style for the report under Style. There are options to add and customise headers and footers to the document. Choose Begin with to add to the start of the report and End with to add to the end of a report. Select the options then either click on Make Report or the Report tab heading. An html report will be generated and saved into the folder containing the current structure.
Generating a CIF
Work|Report|Edit CIF Info brings up a text editor enabling the currently known information to be edited. Any edits made in this window will take precedence over any previous values. We recommend that you make all edits to the .cif file here.
Work|Report|Merge-CIFwill merge all CIF information into one .cif file. There is also the option to select to include/exclude HKL/RES in the cif.- If ShelXL is being used to carry out the refinement, under
Work|Refine|Refinement-Settings-Extrait is possible to select the ACTA command which will generate a .cif and .fcf file. Note, any information input under the Report section will be added to the CIF.
Validating the CIF
Direct access to the IUCr CheckCif is available through Olex2 under Work|Report|CheckCif-Report with the option to include the .fcf (Send FCF) in the submission or not. The report is saved in either html or pdf format into the folder containing the current crystal structure.
It is important to check the .cif file for any errors or problems with the crystal structure. To this end, the IUCr offers a free automated checking service for .cif files at http://checkcif.iucr.org/. To obtain a full report, the structure factors have to be included (Send FCF). Alerts are given under codes ranging from A-G and each has a section title that, if generated directly through the program on the web, can be clicked on for more details of what an alert means. Alert A’s are the most serious and generally need attention while Alert G is probably only advisory. All Alerts that can be solved should be fixed and any more major problems that can’t be fixed should be explained.
Submitting a structure to the CCDC
A CCDC number ready for a paper can be obtained by clicking on the link CCDC Number. You can specify whether this submission is intended for a specific journal, or whether it should be treated as a private communication. In either case, you will receive an e-mail with the CCDC number a few days after your submission. The CCDC is always very happy to receive new structure reports of structures that already exist in the CSD. So next time your structure turns out to be “already in the CSD”, why not submit it as a private communication? It is not necessary to devise a naming scheme and/or prepare any publication material, so it is easily done.
Updating the Chemical Composition
This should be done upon completion of the structure refinement. If the currently assigned chemical composition is different to that displayed on the screen under Work|Toolbox Work the atom types will not appear green. To update the chemical composition:
- Under
Work|Toolbox-Workclick on Click Refine to update the formula.
Click Refine to update the formula. - Under
Work|Solve|Chemical-Compositionsimply enter the formula.
Refinement Data Plots
We have covered reflection data plots before. Here are the remaining plots that are only relevant once you have got a model. These are currently (wrongly) located under Info|Reflection Statistics:
Fo-Fc
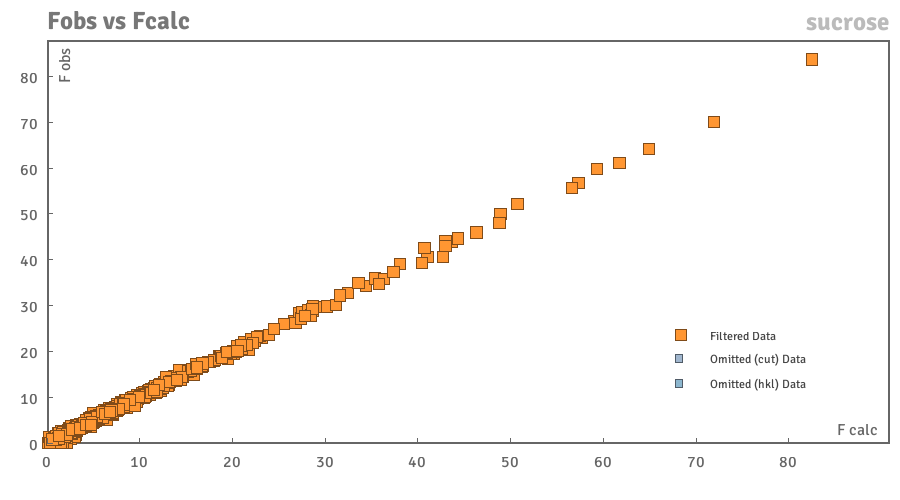
This should be a straight line with Fo-Fc, i.e. the gradient of the line should be ~1 and the intercept ~ 0. Any omitted data are shown in grey. If a reflection appears to be an outlier hovering over it gives relevant information in the following format (Fo, Fc)(h, k, l).
Fo/Fc
Plotted against resolution, should be around 1.0.
Scale factor vs resolution
These should be approximately constant around 1.0 across the whole data range, a low value at high values of 2Theta can indicate that the data is weak or not present in that region.
R1 factor vs resolution
This value will increase for higher angle data.